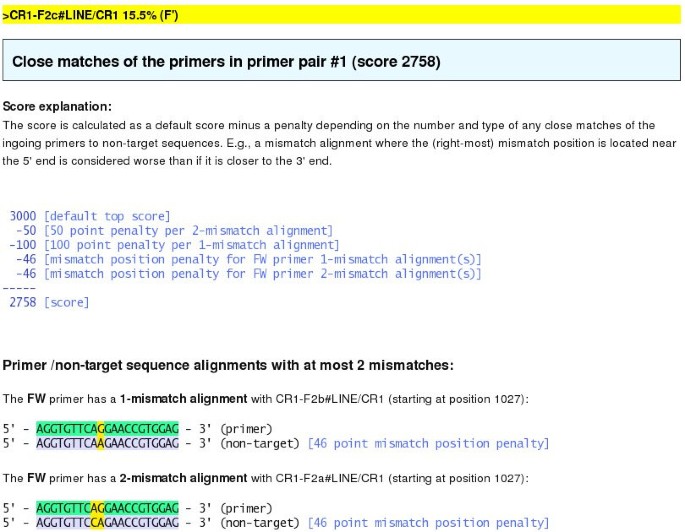
Primique: automatic design of specific PCR primers for each sequence in a family | BMC Bioinformatics | Full Text
Deconstructing the Polymerase Chain Reaction: Understanding and Correcting Bias Associated with Primer Degeneracies and Primer-Template Mismatches | PLOS ONE

A mismatch-tolerant RT-quantitative PCR: application to broad-spectrum detection of respiratory syncytial virus | BioTechniques

Sequential development of several RT‐qPCR tests using LNA nucleotides and dual probe technology to differentiate SARS‐CoV‐2 from influenza A and B - Radvánszka - 2022 - Microbial Biotechnology - Wiley Online Library

Pathogens | Free Full-Text | GoPrime: Development of an In Silico Framework to Predict the Performance of Real-Time PCR Primers and Probes Using Foot-and-Mouth Disease Virus as a Model
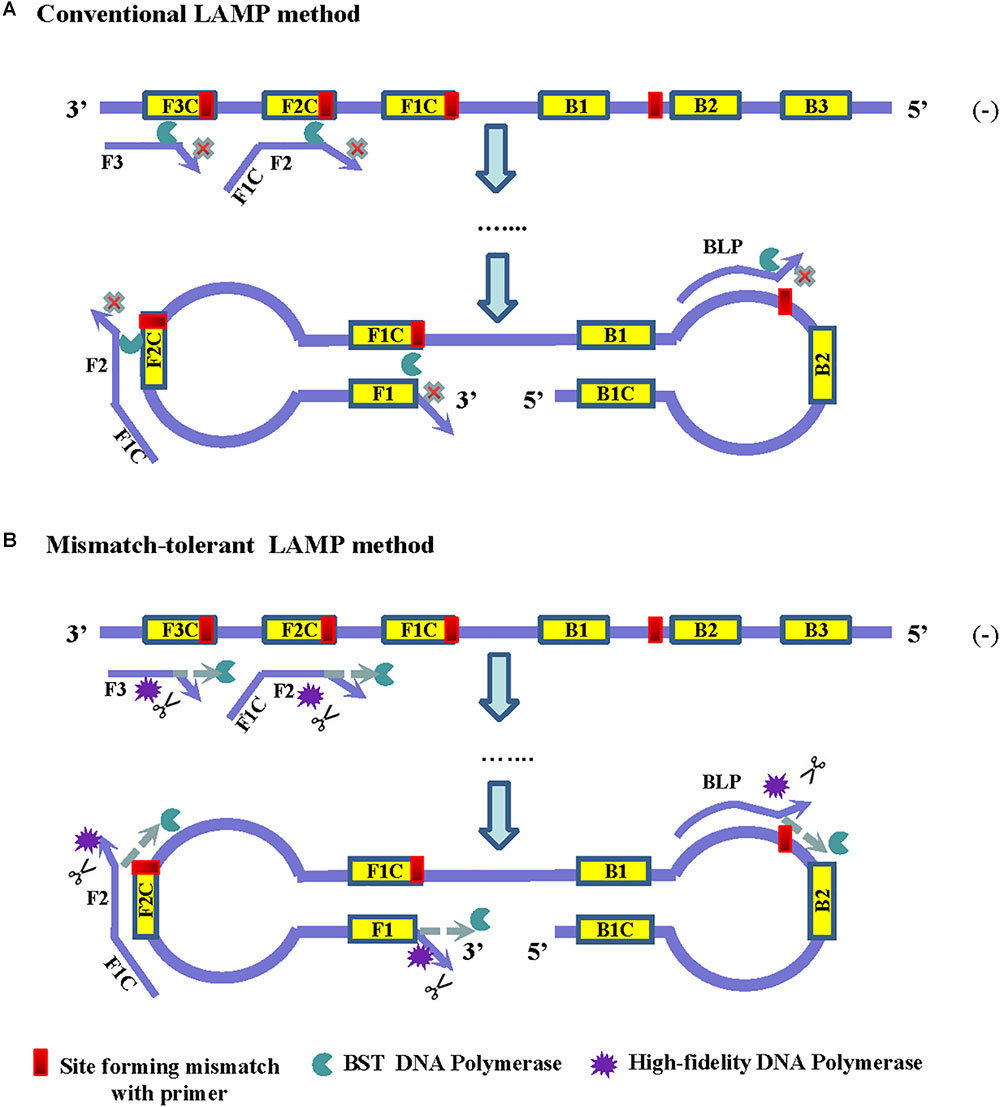
Frontiers | A Mismatch-Tolerant Reverse Transcription Loop-Mediated Isothermal Amplification Method and Its Application on Simultaneous Detection of All Four Serotype of Dengue Viruses
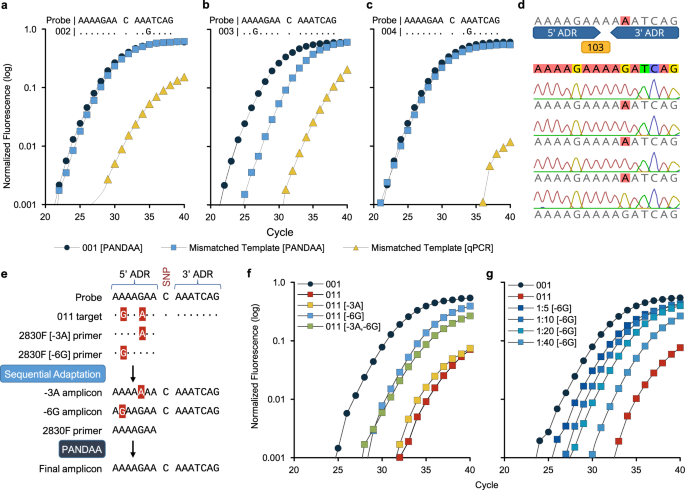
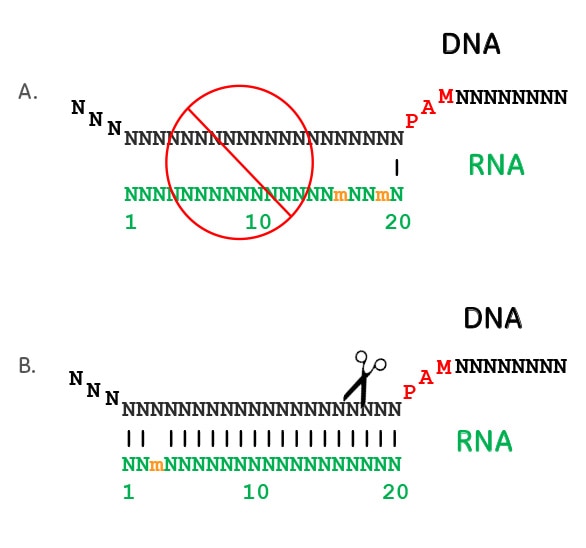
PANDAA intentionally violates conventional qPCR design to enable durable, mismatch-agnostic detection of highly polymorphic pathogens | Communications Biology

PANDAA intentionally violates conventional qPCR design to enable durable, mismatch-agnostic detection of highly polymorphic pathogens | Communications Biology

Profiling the Mismatch Tolerance of Argonaute 2 through Deep Sequencing of Sliced Polymorphic Viral RNAs - ScienceDirect

Pathogens | Free Full-Text | GoPrime: Development of an In Silico Framework to Predict the Performance of Real-Time PCR Primers and Probes Using Foot-and-Mouth Disease Virus as a Model
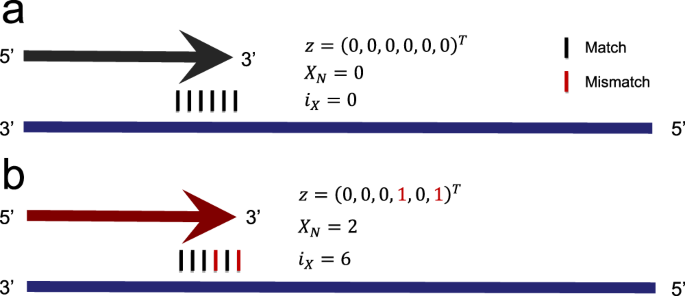
Modeling the Amplification of Immunoglobulins through Machine Learning on Sequence-Specific Features | Scientific Reports